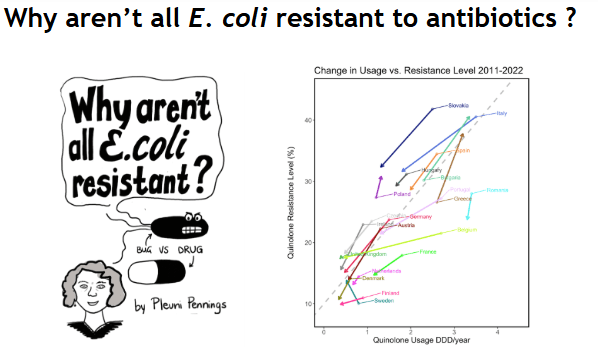
Why aren't all E. coli resistant to antibiotics?
(Seminar in English)
Pleuni Pennings
Department of Biology, San Francisco State University, San Francisco, USA
pennings@sfsu.edu
Friday, March 28
CEFE – Large meeting room – 11:30 a.m.
1919 Route de Mende, Montpellier
The seminar will also be streamed live online.
Link to seminar:https://umontpellier-fr.zoom.us/webinar/register/WN_zgLs6hEhTxSlPV2sPbMsRQ
Access to campus (register before 11 a.m. on SEEM day):https://duo.dr13.cnrs.fr/public/evenement/index
Abstract
We don't know why drug-resistant strains and susceptible strains often coexist and what determines the level of resistance in a pathogen population (e.g.,E. coli,S. pneumonia, HIV). However, there is a lot wedoknow: we know that resistance levels are often stable over years, we know that resistance levels depend on drug usage at the country level, and we know that resistance is often due to many independent acquisition events (whether mutational or through horizontal gene transfer). Here I present a simple population-genetic model, akin to mutation-selection balance which can explain all these observations.